Skin dose maps¶
Functionality that is available¶
- Skin dose map data is calculated to the surface of a simple geometric phantom using the in-built openSkin routines (3D phantom)
- The calculated doses include kVp-dependent backscatter factors and account for any copper filters using data from this paper for each irradiation event; aluminium or other filters are not considered. Where more than one kVp is stored for an irradiation event the mean kVp is calculated, excluding any zero values.
- The phantom dimensions are calculated from the height and mass of the patient; defaults of 1.786 m and 73.2 kg are used when patient height and mass are not available
- Data can be calculated on import to OpenREM, or on demand when a study is viewed. Calculating the skin dose map can take several minutes, so calculating on import may tie up your server for that time, depending on how you have configured the Celery task queue
- Data is recalculated automatically if the patient height or mass stored in the database differs from the values stored in the skin dose map data file. This is useful when patient size information has been imported in to OpenREM after the initial skin dose map data has been calculated
- 3D skin dose map data is shown graphically as a 2D image and a 3D model
- The 3D model can be manipulated using a touch screen
- The user can change the maximum and minimum displayed dose; alternatively, window level and width can be adjusted
- A colour dose scale is shown with a selection of colour schemes
- The skin dose map section can be displayed full-screen
- The calculated peak skin dose, phantom dimensions, patient height, mass and orientation used for the calculations are shown in the top left hand corner of the skin dose map
- If skin dose map display is disabled then fluoroscopy study data can be exported in a format suitable for the stand-alone openSkin routines
The phantom consists of a cuboid with one semi-cylinder on each side (see 3D phantom section of phantom design on the openSkin website for details).
2D visualisation of the 3D data¶
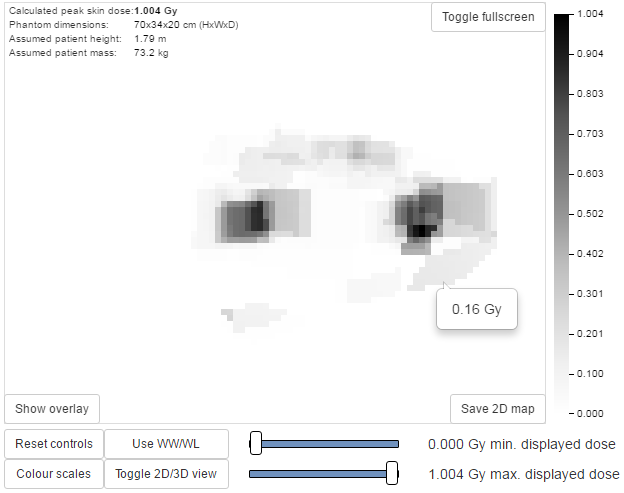
This is a 2D view of the whole surface of the 3D phantom, as though the phantom surface has been peeled off and laid out flat (figure 1). The 2D visualisation includes the following features:
- The skin dose at the mouse pointer is shown as a tool-tip
- Moving the mouse whilst holding down the left-hand mouse button changes the window level and width of the displayed skin dose map
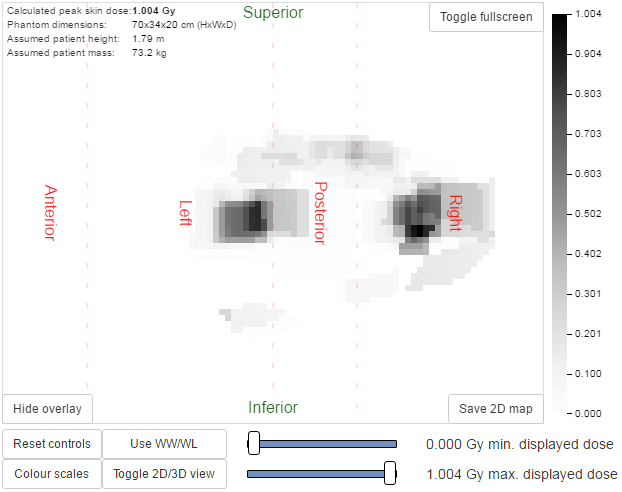
- An overlay indicating the phantom regions and orientation can be toggled on and off. This indicates the phantom anterior, left, posterior and right sides, and also shows the superior and inferior ends (figure 2)
- The current view can be saved as a png file
3D visualisation¶
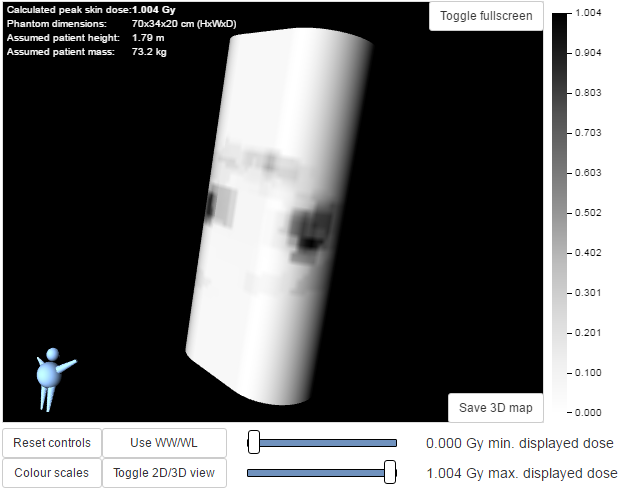
This is a 3D view of the phantom that was used for the calculations, with the skin dose map overlaid onto the surface. The 3D visualisation includes the following features:
- Moving the mouse whilst holding down the left-hand mouse button rotates the 3D model
- Using the mouse wheel zooms in and out
- A simple 3D model of a person is displayed in the bottom left corner. This is to enable the viewer to orientate themselves when viewing the 3D skin dose map
- The current view can be saved as a png file
Skin dose map settings¶
There are two skin dose map options that can be set by an OpenREM
administrator via the Skin dose map settings option in the Config menu:
- Enable skin dose maps
- Calculate skin dose maps on import
The first of these sets whether skin dose map data is calculated, and also switches the display of skin dose maps on or off. The second option controls whether the skin dose map data is calculated at the point when a new study is imported into OpenREM, or calculated when a user first views the details of that particular study in the OpenREM interface.
When skin dose maps are enabled:
- When a user views the details of a fluoroscopy study OpenREM looks for a skin
dose map pickle file on the OpenREM server in the
skin_mapssubfolder ofMEDIA_ROOTthat corresponds to the study being viewed. If found, the skin dose map data in the pickle file is loaded and displayed. Theskin_mapsfolder is created if it does not exist - If a pickle file is not found then OpenREM calculates skin dose map data.
These calculations can take some time. They are carried out in the
background: an animated graphic is shown during the calculations. On
successful calculation of the data the skin dose map is displayed. A pickle
file containing the data is saved in the server’s
skin_mapssubfolder ofMEDIA_ROOT. The file name is of the formskin_map_XXXX.p, whereXXXXis the database primary key of the study - For subsequent views of the same study the data in the pickle file is loaded, rather than re-calculating the data, making the display of the skin dose map much quicker
When calculation on import is enabled:
- OpenREM calculates the skin dose map data for a fluoroscopy study as soon as it arrives in the system
- A pickle file containing the data is saved in the
skin_mapssubfolder ofMEDIA_ROOT - Users viewing the details of a study won’t have to wait for the skin dose map data to be calculated
Exporting data to openSkin¶
If skin dose maps are disabled, and the user has export rights, the user is presented with the option of exporting the study data as a csv file that is formatted for use with a stand-alone installation of openSkin. The user must be in the detail view of the study they wish to create the exposure incidence map for, and then click on the link to create the OpenSkin export (figure 4).
Instructions for openSkin¶
Download the openSkin repository as a zip file from openSkin downloads. To use openSkin as a stand-alone application you need python 2.x and the pypng python library.
- Extract the contents of the zip file into a folder on your computer and run python main.py from a command line and answer each question.
- See phantom design for details of the 2D and 3D phantoms.
- When asked for the source csv file use the one exported from OpenREM
- Depending on the number of events in the export and the power of your computer the calculations can take a few minutes
Two files will be produced - a textfile called skin_dose_results.txt and a
small image called skin_dose_map.png
Results text file¶
It should look something like this:
File created : 04/21/15 17:42:45
Data file : C:/Users/[...]/exports-2015-04-21-OpenSkinExport20150421-162805246134.csv
Phantom : 90.0x70.0 3d phantom
Peak dose (Gy) : 0.50844405521
Cells > 3 Gy : 0
Cells > 5 Gy : 0
Cells > 10 Gy : 0
The peak dose is the peak incident dose delivered to any one-cm-square area. If any of these 1 cm2 areas (referred to as cells) are above 3 Gy, then the number of cells in this category, or the two higher dose categories, are listed in the table accordingly.
Incidence map image file¶
The image file will be a small 70x90 px PNG image if you used the 3D phantom, or 150 x 50 px PNG if you used the 2D phantom. With both, the head end of the table is on the left.
The image is scaled so that black is 0 Gy and white is 10 Gy. For most studies, this results in an incidence map that is largely black. However, if you use GIMP or ImageJ or similar to increase the contrast, you will find that the required map is there.
A native and ‘colour equalised’ version of the same export are shown below:
Limitations¶
Skin dose map calculations do not currently work for all systems. Siemens Artis Zee data is known to work. If skin dose maps do not work for your systems then please let us know via the OpenREM Google Group.
openSkin is yet to be validated independently - if this is something you want to do, please do go ahead and feed back your findings to Jonathan Cole at jacole.